Scientists unearth a clue to the molecular mechanisms involved in N2O reduction by deep-sea hydrothermal vent bacteria.
Nitrous oxide (N2O) is the third most potent greenhouse gas after carbon dioxide and methane. It can also be oxidized by physical processes to form ozone-depleting substances. Atmospheric concentrations of N2O have increased since the preindustrial era, making N2O reduction a global challenge.
The only known biological sink of N2O in the biosphere is microbial denitrification. Denitrification is a series of reduction reactions starting with nitrate and ending with the reduction of N2O to nitrogen gas, with no greenhouse effect. This reaction is unique to microorganisms possessing N2O reductase (N2OR; NosZ), highlighting the importance of identifying the molecular mechanisms mediating high N2O reduction activity.
Researchers at Hokkaido University, in collaboration with colleagues at the Institute of Physical and Chemical Research (RIKEN) and the University of Washington, investigated the molecular mechanisms underlying N2O reduction of a microbial species, Nitrosophilus labii HRV44T, which had been discovered by Hokkaido University researchers in 2020, from a deep-sea hydrothermal vent. The team recently published their results in the journal iScience.
Nitrosophilus labii HRV44T is a thermophilic chemolithoautotroph isolated from a deep-sea hydrothermal vent in the Okinawa Trough, Japan. It grows using hydrogen as an electron donor and N2O as an electron acceptor. (Photo by Muneyuki Fukushi, Hokkaido University)
The research team developed a method that enabled them to analyze time-series gene expression at a genome-wide level, called the transcriptome, using RNA extracted from very few cells.
"Time series transcriptomic analysis of HRV44T in response to N2O was more challenging than expected," said corresponding author Sayaka Mino, Assistant Professor at the Faculty of Fisheries Sciences, Hokkaido University. "We have performed transcriptomic analysis using methods often used in microbial studies, but we failed to capture the gene expression dynamics over short time scales because we could not get enough RNA from just a few cells. The method demonstrated in the current study requires only 1 ng of messenger RNA (mRNA), making it useful for analysis at low cell densities, from which RNA extraction is difficult."
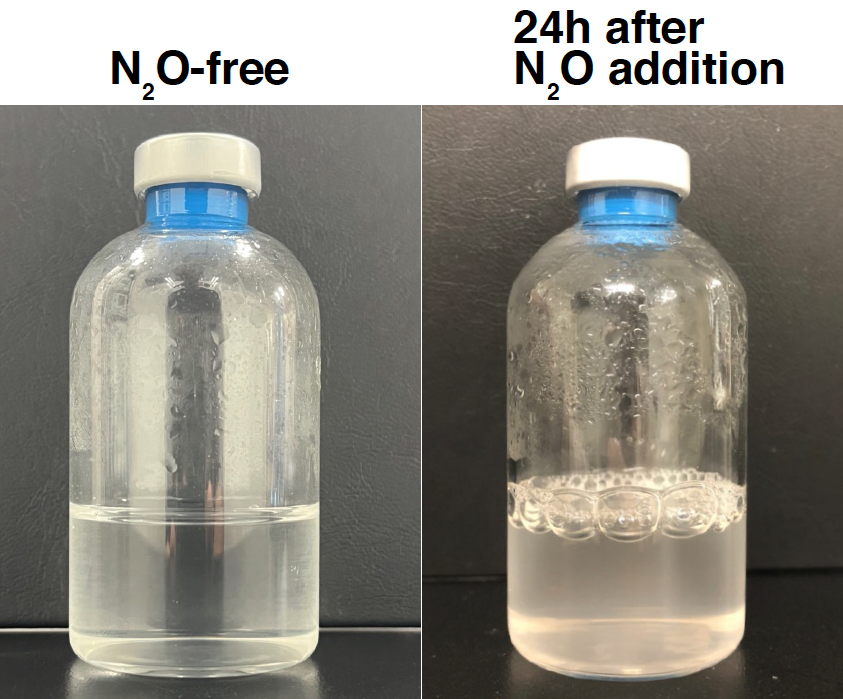

Strain HRV44T rapidly respires N2O, forming bubbles at the gas-liquid interface. (Photo by Jiro Tsuchiya, Hokkaido University)
The time series transcriptomic profiling of HRV44T demonstrated that N2O is not a critical inducer of denitrification gene expression, including nos genes, which are expressed under anaerobic conditions even in the absence of nitrogen oxides as electron acceptors.
"We hypothesize that this feature may contribute to efficient energy metabolisms in deep-sea hydrothermal environments where alternative electron acceptors are occasionally depleted", said Robert M. Morris, Associate Professor at the University of Washington.
Jiro Tsuchiya, the first author and a JSPS research fellow DC2 at Hokkaido University, and colleagues conducted a statistical analysis of time series data. "Our findings suggest that the denitrification gene nosZ is negatively regulated by transcriptional regulators that typically function as transcriptional activators in response to environmental changes. Although we still need to investigate this result, our study extends the understanding of the regulatory mechanisms controlling gene expression in N2O-reducers and may help increase their ability to respire N2O", said Tsuchiya.
Deep-sea hydrothermal environments have steep chemical and physical gradients, making them hotspots for bioresources. This study demonstrates the potential for microorganisms in these environments to contribute to N2O mitigations that may help combat climate change. Search for microbial resources with high greenhouse gas reduction efficiency, optimization of their abilities, and elucidation of molecular mechanisms specific to these microorganisms will contribute to developing technologies for environmental remediation by microorganisms.
Original article:
Jiro Tsuchiya, Sayaka Mino, Fuki Fujiwara, Nao Okuma, Yasunori Ichihashi, Robert M. Morris, Brook L. Nunn, Emma Timmins-Schiffman, Tomoo Sawabe. Time course transcriptomic profiling suggests Crp/Fnr transcriptional regulation of nosZ gene in a N2O-reducing thermophile. iScience. October 22, 2024.
DOI: 10.1016/j.isci.2024.111074
Funding:
This work was partially supported by the Institute for Fermentation, Osaka; Takahashi Industrial and Economic Research Foundation; and, Japan Society for the Promotion of Science (JSPS) KAKENHI (17K15301, 21KK14913), JSPS Overseas Research Fellowships and JSPS Research Fellowships for Young Scientists.